Ribonucleotide Reductase
Nucleic Acid Metabolism
Ribonucleotide reductase reduces ribonucleotides to deoxyribonucleotides, a crucial step in DNA synthesis.
Ribonucleotide reductase catalyzes the reduction of ribonucleotides to deoxyribonucleotides, the essential building blocks for DNA synthesis. This enzyme is critical for the production of deoxyribonucleotides (dNTPs), which are required for DNA replication and repair. RNR specifically reduces the 2’-hydroxyl group of ribonucleotides (ATP, GTP, CTP, and UTP) to the corresponding deoxyribonucleotides (dATP, dGTP, dCTP, and dTTP). This reduction is coupled with the oxidation of a thiol group, typically on a cysteine residue within the enzyme’s active site, and is essential for maintaining an adequate supply of dNTPs in cells.
The enzyme operates through a complex mechanism involving radical chemistry. Ribonucleotide reductase consists of two main subunits: the R1 subunit, which contains the catalytic site, and the R2 subunit, which contains a radical-generating tyrosyl radical that is crucial for the reduction process. The reaction also requires electron transfer from a thioredoxin or glutaredoxin system, which provides the reducing equivalents needed to convert ribonucleotides into deoxyribonucleotides.
The activity of ribonucleotide reductase is tightly regulated to ensure the balanced production of the four dNTPs. This is critical because an imbalance in the levels of dNTPs can lead to DNA replication errors and genomic instability. The enzyme’s activity is regulated both at the transcriptional level and through allosteric regulation. The availability of dATP and dTTP acts as a negative feedback mechanism to control the enzyme’s activity, while ATP and dGTP activate it, promoting balanced nucleotide production.
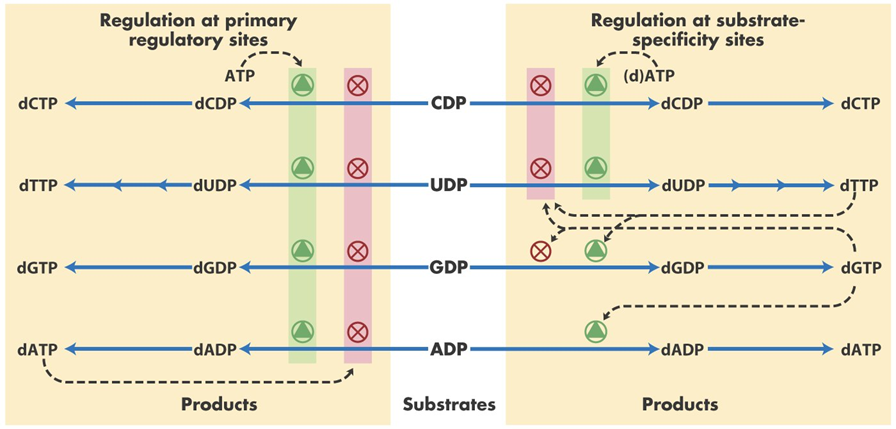
Ribonucleotide reductase is tightly regulated to ensure a balanced supply of deoxyribonucleotides, preventing both shortages and excesses that could disrupt DNA synthesis. The enzyme is regulated by allosteric effectors, which control the relative production of each dNTP. For instance, dATP serves as a feedback inhibitor, reducing the enzyme’s activity to avoid overproduction of deoxyribonucleotides. On the other hand, ATP and dGTP activate the enzyme to stimulate dNTP production, particularly when the cell is preparing for DNA replication.
The regulation of RNR is also linked to the cell cycle. During DNA replication, the demand for deoxyribonucleotides is heightened, so the activity of ribonucleotide reductase is upregulated to meet these needs. Conversely, in the resting or quiescent phase, the enzyme’s activity is downregulated, conserving resources and preventing excessive dNTP synthesis. In some cells, p53, a tumor suppressor protein, can also influence RNR activity by inhibiting its function, particularly in response to DNA damage or stress, thus preventing the replication of damaged DNA.
In addition to allosteric and cell cycle regulation, post-translational modifications such as phosphorylation can influence the activity of ribonucleotide reductase, further fine-tuning its function during different cellular conditions. Through these regulatory mechanisms, ribonucleotide reductase ensures that dNTPs are produced in the correct amounts, supporting accurate DNA replication, repair, and overall genomic stability.


